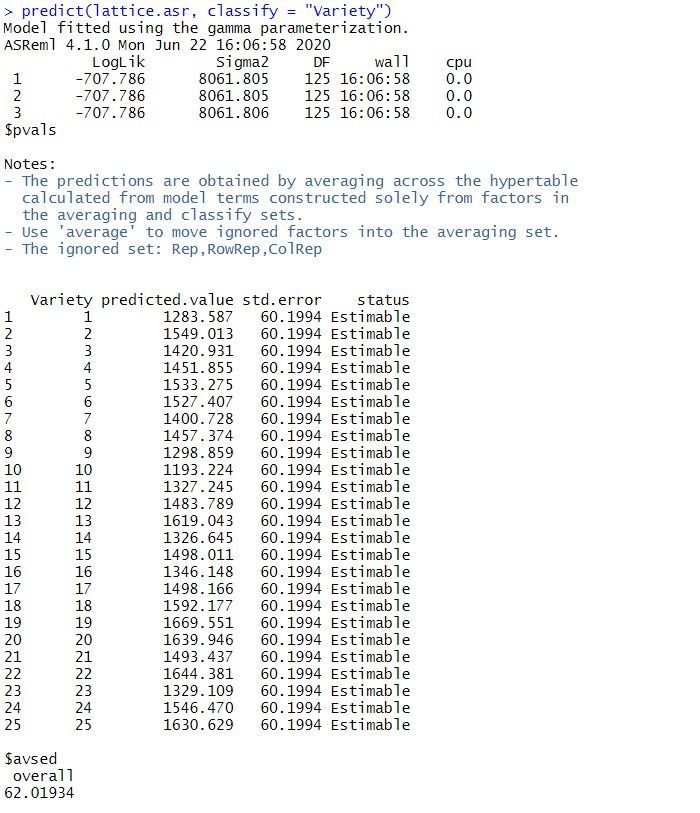
Heritabilities of traits were estimated using BLUP (Best Linear Unbiased Prediction) evaluation techniques. Recording of body measurements was continued for 3 years since 2010 to 2012. *Corresponding author: paper was presented in International Buffalo Congress 2019, February 18-20, Lahore, PakistanĬurrent study was conducted to find out heritability estimates of some body measurements of Nili Ravi buffaloes maintained at 5 Livestock Experiment Stations in Punjab Pakistan.

Kuthu 5ġDepartment of Livestock and Poultry Production, Faculty of Veterinary Sciences, Bahauddin Zakariya University, Multan, Pakistan 2Department of Breeding and Genetics, Faculty of Animal Production and Technology, Cholistan University of Veterinary and Animal Sciences, Bahawalpur- 63100, Pakistan 3Buffalo Research Institute, Pattoki-55300, (Punjab), Pakistan 4Department of Theriogenology, Faculty of Veterinary and Animal Sciences, Gomal University, Dera Ismail Khan-29220, Pakistan 5Faculty of Veterinary Sciences, University of Poonch, Rawalakot 0.809, a 1.5% relative difference) I don't know if that is big enough to be of concern or not.Content for New Div Tag Goes Here HERITABILITY ESTIMATES OF SOME BODY MEASUREMENTS IN NILI RAVI BUFFALOES The H^2 estimate looks right on, but the H^2c estimate is a little different (0.797 vs. I'm going to take the means of these values ( not double the values, since these are already the variances of differences) vblue <- mean(vv2)
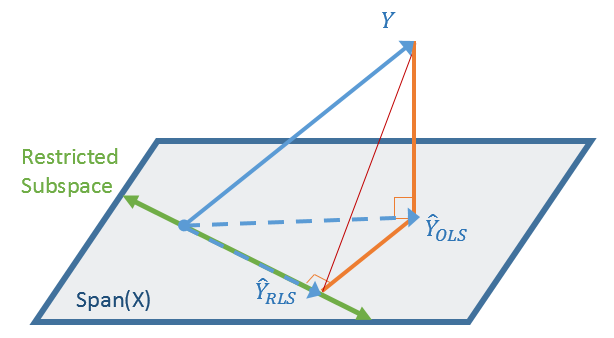
vblup <- 2*mean(vv1)įor the model with gen fitted as a fixed effect, let's extract the variances of the parameters relating to genotypes (which are differences in the expected value from the first level): vv2 <- diag(vcov(model2)) If these were all the same then the mean variance of a difference of any two would be just twice the variance (Var(x-y) = Var(x)+Var(y) by construction the BLUPs are all independent). The relative variation is approximately 5.4e-4. I don't know whether they should all be identical (and this is a roundoff issue) (uv <- unique(vv1)) They're not all the same, but they're pretty close. This is a 1x1x24 array (effectively just a vector of variances we could collapse using c() if we needed to). Get conditional variances of BLUPs: rr1 <- ranef(model1,condVar=TRUE) Model1 <- lmer(yield ~ 1 + rep + (1|gen) + (1|rep:block), john.alpha) I'm not that familiar with this variance partitioning stuff, but I'll take a shot.
#LEAST SQUARE MEANS ASREML HOW TO#
How to compute the vblup and vblue (mean variance of difference) in lme4 equivalent to predict()$avsed ^ 2 of asreml ? Model2 <- lmer(yield ~ 1 + gen + rep + (1|rep:block), dat) Varcomp <- ame(print(varcomp, comp = "Variance")) I am trying to replicate the above using the R package lme4. Vblue <- predict(model2, classify="gen")$avsed ^ 2 # mean variance of a difference of two adjusted treatment means (BLUE) Model2 <- asreml(yield ~ 1 + gen + rep, data=dat, random = ~ rep:block) Vblup <- predict(model1, classify="gen")$avsed ^ 2 # mean variance of a difference of two BLUPs Model1 <- asreml(yield ~ 1 + rep, data=dat, random=~ gen + rep:block)
#LEAST SQUARE MEANS ASREML CODE#
Given below is the code for analysis of a resolvable alpha design (alpha lattice design) using the R package asreml.
